Overview
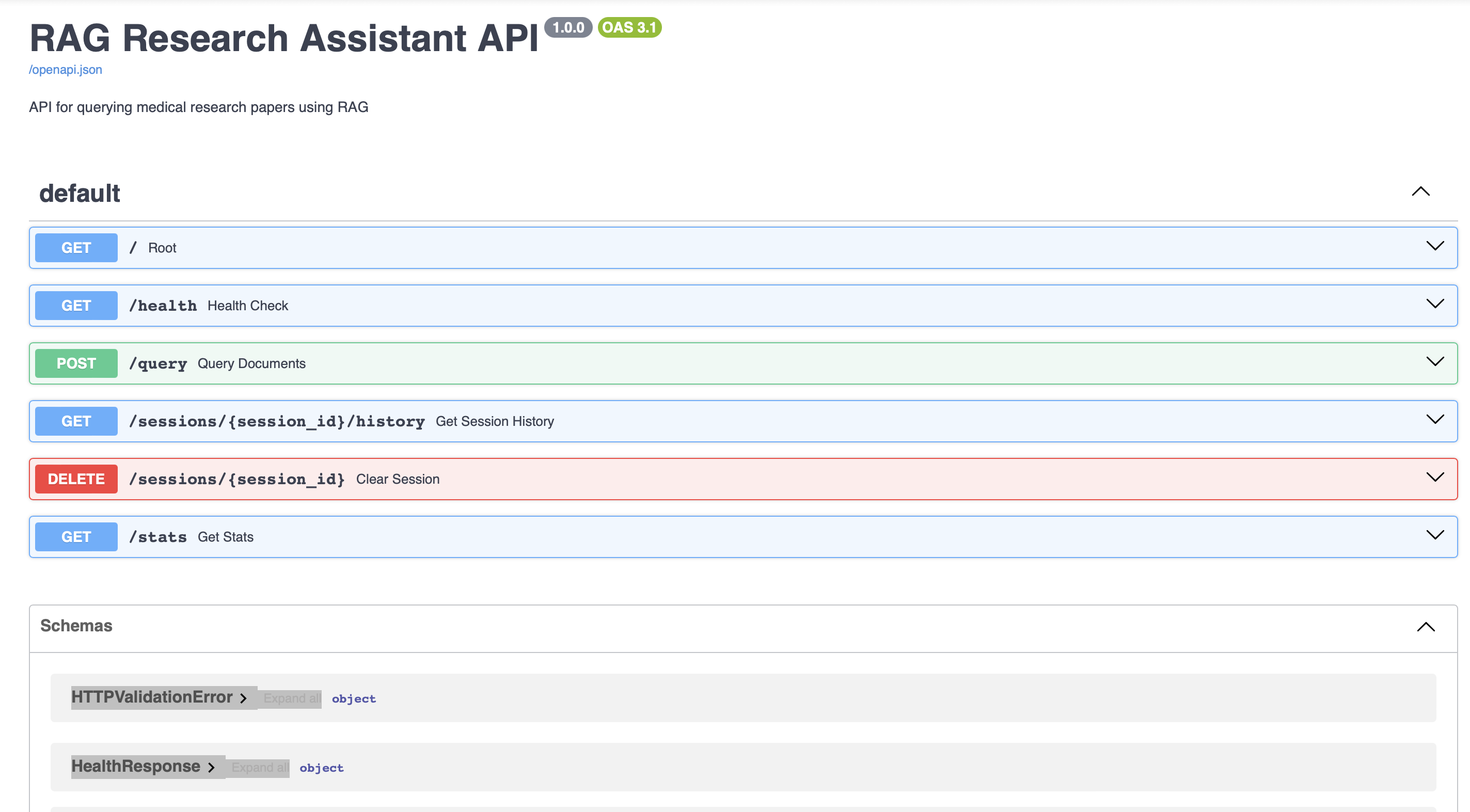
An end-to-end RAG pipeline that enables natural language querying of peer-reviewed medical research papers on brain tumor detection. Users ask questions and receive accurate, citation-grounded answers sourced directly from the indexed PDFs, delivered through a FastAPI backend and Streamlit frontend.
The idea came out of a Spring 2025 master’s-level project on deep learning for brain tumor detection from MRI images, which pushed me to think less about individual models and more about supporting the research workflow itself.

Dataset & Scale
| Metric | Value |
|---|---|
| Research papers indexed | 20 papers (478 pages) |
| Text chunks indexed | 2,184 |
| Average response time | 2–5 seconds |
Features
- Natural language question answering over medical PDFs
- Citation-grounded responses sourced directly from indexed papers
- Session-based conversational memory
- Async REST API with multi-user support
- Interactive Streamlit web interface
Architecture
PDF Papers → PyPDF Parsing → Text Chunking → OpenAI Embeddings
↓
User Query → Streamlit → FastAPI → ChromaDB Retrieval → GPT-4o-mini → Response + Citations
Tech Stack
Python FastAPI Uvicorn OpenAI API (GPT-4o-mini) LangChain ChromaDB Streamlit PyPDF Retrieval-Augmented Generation
Note: This project is intended for research and educational use only and is not suitable for clinical or diagnostic decision-making.